Note: This post was updated on 30 July 2022 with new information.
Recently, I suggested to a person on one of the many genetic genealogy
forums that you may be able to use Y-DNA, mtDNA, and X-DNA to narrow down a
particular line of descent for an individual autosomal match. I was immediately rebuffed
by someone who basically said the following, “Y-DNA, mtDNA, and
X-DNA have no place in an autosomal DNA study.”
I responded, “While the
inheritance patterns of other types of DNA are different, they still can be used (in some
cases) to determine possible lines of relationship.” The responder was not
convinced and was agitated that I would even suggest such tactics. However, a
multidisciplinary approach to DNA may aid in eliminating certain lineages from
consideration when determining the origin of a match.
I thought most folks who have dipped their toes into the genetic genealogical pool would have known this; however, after being involved in several discussions on this topic, I felt the need to author several posts in this regard.
The Larger Study
In the past, I have used autosomal and Y-DNA to determine
relationships, but up until recently, I have not had the opportunity to use
mtDNA and X-DNA to do the same. The examples found in this discourse are
part of an ongoing study that uses autosomal DNA, X-DNA, and mtDNA from nearly 30
participants who are related to our ancestral couple: Ancestral Father (AF)
and Ancestral Mother (AM).
In trying to determine a particular genealogical problem of
whether a person was the natural or adopted child of one of AF and AM’s
children, I used autosomal and mtDNA to test a null hypothesis. The null hypothesis (names changed) was similar to the following: H0: Mary Smith was not the natural daughter of Jane Doe. By comparing
the autosomal DNA from descendants of AF and AM and matching mtDNA transmitted from
AM, the null hypothesis was rejected. Descent in this case was verified.
While this study is ongoing and we are still amassing evidence in hopes of submitting the finished product for publication, the surname of the family and all
additional specific information concerning these lineages (including references
and location) have been omitted to protect the original study.
This has been
done so that my work is not co-opted by another individual who has been
known to plunder my research on this family and publish it online. I began this study during 2014 when I first became aware of several issues related to this lineage. This project involves a surreptitious narrative that is far more involved than the DNA analysis.
This isn't an analysis of this particular study per se; it is an exercise to show that using other DNA types can aid an autosomal study. So while the names, locations, and documentation of sources are absent, the techniques are present and that's what is important to this post.
The Current Problem
While working on the DNA analysis, we have identified two unknown
individuals genetically related to this family: U1 –
an adopted individual and U2 – who cannot be placed as of yet. It is
believed that U2 descends from one of the Ancestral Mother’s siblings. Until
more information is available, it is uncertain how U2 is connected to the family.
This tangential part of the study was conducted to aid U1 in identifying her
birth mother. Therefore, our task is to narrow the possible lines of descent
for U1’s birth mother that may possibly lead to her identity.
Background of the Family
While AF and AM had 11 children, three failed to attain the
age of majority and their names are currently unknown. These additional children were listed in the number of children born to the mother in both the 1900 and 1910 censuses. Seven of 11 children were listed as living in the 1900 census and six of 11 were numbered as living in the 1910. E-Prime (E0) died before 1900 and her sister A-Prime (A0) expired prior to 1910.
Of the eight
children who attained majority, one son never married nor had any known issue. Descendants of six of the remaining seven children have tested. We
are still attempting to attract descendants of the youngest child S-Prime (S0)
to test. While S0 had two sons and two daughters, neither of her daughters produced
issue. Twenty of AF and AM's 35 grandchildren produced issue; descendants of 14 grandchildren are current participants. We are still soliciting participants to strengthen our original study's results.
The known children of the ancestral parents are as follows:
- E-Prime (E0) – daughter, born 1856; 8
participants
- K-Prime (K0) – daughter, born 1858; 2
participants
- J-Prime (J0) – son, born 1862; 9 participants
- Son, born 1864 (no issue)
- A-Prime (A0) – daughter, born 1867; 3
participants
- W-Prime (W0) – son, born 1869; 3 participants
- F-Prime (F0) – son, born 1871; 2 participants
- S-Prime (S0) – daughter, born 1875; no known
tested descendants
Additionally, there are several others in this project
- Six descendants from AF’s surname lineage: X1-X6, with four descended from AF’s siblings.
- Two likely descendants of J0 (not in the data
presented). These two share sizeable amounts of autosomal DNA with J0’s
descendants as well as to descendants of J0’s wife’s sisters.
- Two unknowns: U1 and U2.
It is also noted that all the descendants of E0 and K0
share double relationships. Additionally, the grandchildren (E1, E2,
& E3) of one of E0’s sons share double relationships with all the descendants of J0. Finally, the descendants of the oldest daughter of W0 share double relationships with the descendants of the oldest granddaughter of A0. None of
these double relationships impact the results of this present analysis.
With U1, it is possible narrow down her birth relationship
to two possible lines, but we are still soliciting family members to test to
determine her mother’s exact identity. In this discourse, I’ll show how we are
using autosomal, X-DNA, and mtDNA in an attempt to narrow the possibilities.
Autosomal DNA
In looking at the triangulated segments between U1 and
others represented in this project, it is obvious that U1 is related to, if not
descended from, AF and AM. The triangulated segments are listed below.
We have
also separated the triangulated segments between U1 and only those descended
from A0. We’ll elaborate on this in the next paragraph. I have also separated
out a triangulated segment between U1 and two descendants of a sister of AF. This is somewhat inconclusive, as the relationship could come through one of U1’s other lineages.
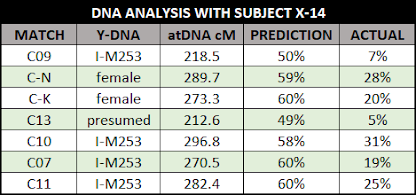
The amount of DNA U1 matches with various tested individuals
narrows to one child of AF & AM who is the likely ancestor of U1. In the
chart below, except for the matches with J1 (at 50.6 cM) and E2 (48.6 cM), the larger matches are
found among the descendants of A0.
J1 is three generations from AF and AM. Her
father was the last born grandchild of AF and AM and places her a generation or
more above all other participants; therefore, she would likely share more than others at a further distance. As far as E2 (a brother of E1 and E3), he generally shares higher amounts of autosomal DNA from this ancestral couple than others descended from his great-grandmother with the exception of a second cousin (E6).
U1’s two largest autosomal DNA shares are with A1 and A3.
Both individuals share 103.7 cM autosomal DNA and a little over 20 cM on
the X-Chromosome. Using the Shared cM Project, the most likely predicted relationship
for both A1 and A3 with U1 is that of second cousins, once removed. A2 is
predicted as a third cousin, once removed. While other relationships are
possible, these better fit the scenario in question. Therefore, it is likely
that U1 descends from A0 – the fifth known child of the ancestral parents.
X-DNA
While the data from our analysis of 17 X-DNA matching
segments among the participants is currently inconclusive, there are some possible theories that could
be further tested. The chart below shows the X-DNA inheritance of 12 tested
known descendants of AF and AM. F0’s living descendants are not present due to a
double male lineage that negates the transmission of X-DNA in the living descendants of F0. It is possible that
some of the untested descendants of one of S0's sons will have additional X-DNA matches. This
remains to be seen.
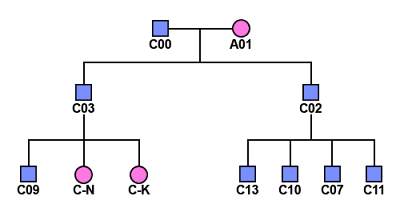
In the above chart, the circles represent females and
squares represent males. The pink circles and squares represent X-DNA only
coming from AM. The purple represents X-DNA possibly inherited through both AF and AM.
The half purple and gray represent X-DNA inheritance that comes from AF, AM, and/or
a shared paternal relative in the descendants of E0 and K0. This is not germane
to this analysis, but it is to the original research project.
The 17 segments are depicted below. Pay particular attention to the eight matching segments with A1 and the three matching segments with U1.
Note: The matches represented are inter-prime families and not intra-prime family. For example, the matches are shown for E5 and E6 with J5; however, the 11.6 cM match between E5 and E6 is not shown.
The matches in black between the descendants of E0 and K0
are probably from a relationship outside of AF and AM’s family. E0’s children
are first cousins to both K0’s daughter and her husband (the son of E0’s
sister-in-law).
Any matches with J0’s and W0’s descendants are from AM and
could come from her mother, father, or a combination of both, but not through
AF. The matches between E0’s descendants an A0’s descendants could come from AF’s
mother, AM’s parents, or any combination thereof.
The longest X-DNA match (in red) is between E5 and A1 at 41.9 cM.
However, A1’s match to E5 has overlapping matches with J3 and J4 (in light blue) at the
beginning of the segment and overlapping matches with W1 and W2 (in dark green) at the end of the
segment. The overlaps would come from one or both of AM’s parents. It is
noteworthy that A1’s matches with this family cover most of the X-chromosome –
only 35-40 cM is not represented by one of her X-chromosomes.
E5’s match with U1 does not correspond with A1’s match with
W1 and W2. This signifies that at least part of E5’s match with U1 comes from
either the other parent of AM or from AF’s mother. This may be the case with part
or all A1’s match with U1. At this juncture, it is impossible to tell, but we
are continuing to look for other possible X-DNA participants.
Since A3's match with U1 is not triangulated with anyone else, this segment could have been inherited through a spouse of one of A0’s descendants and not directly from AF or AM.
At this point, the X-DNA analysis is inconclusive; however, additional X-DNA matches may shed additional light on the connection. What we can conclude is that U1 matches E5, A1, and A3 on the X-chromosome through lines that share the unique transmission of X-DNA.
mtDNA
As mitochondrial DNA is passed from mother to child, we are
fortunate to have three participants that share the same mtDNA haplogroup
(K1a4a1) with their ancestral mother AM. They are E8, K1, and A3. These three
individuals have an unbroken female descent from AM.
Fortunately for this exercise, U1 also shares the K1a4a1 haplogroup. Before we proceed, there is a caveat. Just because there is a matching haplogroup, this is not conclusive by itself as coming directly from AM. It could have been passed through an unrelated ancestor or someone with deeper ancestry to AM’s family.
While this haplogroup could be shared from descendants of
AM’s sisters, we can partially eliminate this possibility. AM had two
sisters reach maturity. One could not be located after 1850. Additionally, her
name is not listed in her mother’s 1879 will. Unfortunately, we cannot rule out
that she lived beyond 1850 and produced issue. This will always be a possibility until her death without issue is confirmed.
The other sister had four daughters and a son. We can rule
out the son, as he could not pass his mtDNA to his children. Of the four daughters,
it does not appear that shared mtDNA was passed to present descendants through any of
these children.
- Daughter One – had one daughter who produced one
son.
- Daughter Two – only had one son.
- Daughter Three – had one daughter who produced
no issue.
- Daughter Four – had a son and an adopted daughter.
This, along with the larger match with A0’s descendants, mtDNA
strengthens the theory that U1 is a female line descendant of A0.
Conclusion
The amount of shared DNA between U1’s and AF & AM's descendants
indicates that U1 is descended from this couple. There is also matching mtDNA
that is consistent with AM’s mtDNA.
Other considerations include
the following inconclusive results:
- U1’s matching to two descendants of AF’s sister (which may or may not come from
other lineages),
- The possibility of inheriting X-DNA from AF’s mother, and
- The
lack of likely mtDNA matches among the known descendants of AM’s traceable sister.
As the amount of autosomal DNA shared with
A1, A2, and A3 indicates descent from A0, let’s look at the possibilities in
A0’s family.
A0 and her husband produced six children – four sons and two
daughters. Three sons died as children. The remaining son (A0b) is the ancestor of
A1 and A2. He can be eliminated, as he could not pass on his mitochondrial DNA. Additionally, A0b produced 11 children who all lived to the age of majority.
The eldest daughter (A0a) had two sons. These sons could not pass their mtDNA to anyone.
This leaves the third child (A0c) – a daughter who was the
great-grandmother of A3.
A0c had nine children – all attained the age of majority.
Five sons can be eliminated.
That leaves four daughters: A0c3, A0c6, A0c7, and A0c8.
A0c6 is A3’s grandmother who had one daughter – A3’s mother.
Any relationship through this lineage would produce a larger amount of shared autosomal DNA. This
line can be eliminated.
A0c3 left the family home prior to 1940 and moved to New York City. She and her husband produced no children.
A0c7 had one daughter who remained in the local area at least through 1999 and may be living there presently. She would have been 27 at the time of U1’s
birth.
A0c8 also had one daughter – she would have been 21 at the
time of U1’s birth. She remained in the same county where U1 was born and where
the adoption occurred. She only moved to a neighboring county as a senior citizen.
In the above possibilities, the relationship of U1 being
born to a daughter of A0c7 or A0c8 would make her a second cousin to A3;
a second cousin, once removed to A1; and a third cousin to A2.
While a share of 103.7 cM is low for second
cousins, the range for relationship according to the Shared cM Project is
41-592 cM. The average share for this relationship is 229 cM; therefore, 103.7 cM is consistent with
other second cousin relationships.
This exercise allows us to narrow down possible individuals
to test. Although this project did not set out to find the birth family of an
adopted child, it has provided us this opportunity. The narrowing down the approximate
ancestry of U1 required us to use genealogical records and a multidisciplinary DNA
approach.
With autosomal DNA pointing to
descent from A0 and without the use of mtDNA, the exercise would have required us to consider numerous lines
in A0’s family. By using mtDNA, we narrowed the possibilities to only two
lines instead of a possible 22 (2 children from A0a, 11 from A0b and 9 from A0c).
I hope this has shown that, by using other forms of DNA in
tandem with autosomal DNA, a multidisciplinary approach may be able to narrow
your search to a smaller number of possible connections.